matRad
An open source dose calculation and treatment planning toolkit for radiotherapy research and education
matRad: An open source dose calculation and treatment planning toolkit for research and education
matRad is an open source software for radiation treatment planning of intensity-modulated photon, proton, and carbon ion therapy started in 2015 in the research group Radiotherapy Optimization within the Division of Medical Physics in Radiation Oncology at the German Cancer Research Center - DKFZ.
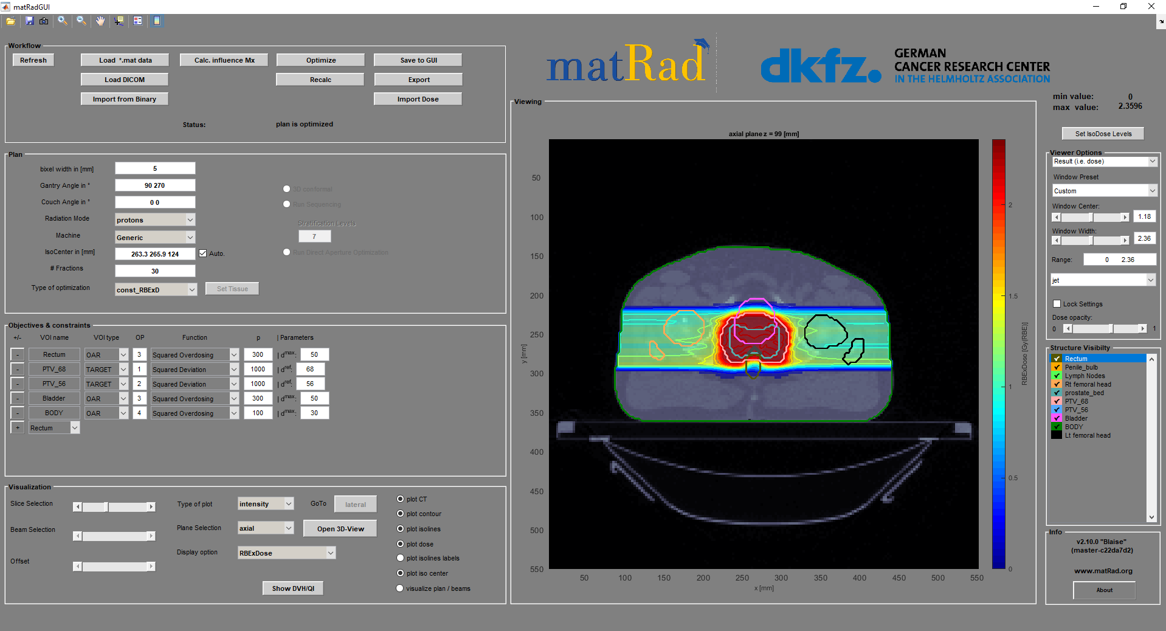
matRad targets education and research in radiotherapy treatment planning, where the software landscape is dominated by proprietary medical software. As of August 2022, matRad had more than 130 forks on GitHub and its development paper was cited more than 160 times (according to Google Scholar). matRad is entirely written in MATLAB and mostly compatible to GNU Octave.
matRad comprises
- MATLAB functions to model the entire treatment planning workflow
- Example patient data from the CORT dataset
- Physical and biological base data for all required computations
In particular we provide functionalities for
- Pencil-beam dose calculation for photon IMRT and proton/carbon IMPT
- Monte Carlo dose calculation for photon IMRT (with ompMC) and proton IMPT (with MCsquare)
- Non-linear constrained treatment plan optimization (based on physical dose, RBE-weighted dose and biological effect) using IPOPT or Matlab’s fmincon
- Multileaf collimator sequencing
- Basic treatment plan visualization and evaluation
- Graphical User Interface
- Standalone Executable (using the Matlab Runtime)
matRad is constantly evolving. If you are interested in working with us or are looking for a special feature do not hesitate and get in touch.
Philosophy
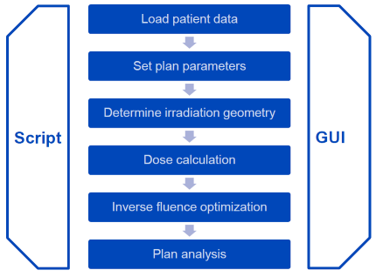
A script to recreate the above treatment plan within matRad 2.10.1 is shown below:
matRad_rc
% load patient data, i.e. ct, voi, cst
load PROSTATE.mat
% meta information for treatment plan
pln.radiationMode = 'carbon'; % either photons / protons / carbon
pln.machine = 'Generic';
pln.numOfFractions = 30;
% beam geometry settings
pln.propStf.bixelWidth = 5; % [mm] / lateral spot spacing for particles
pln.propStf.gantryAngles = [90 270]; % [?]
pln.propStf.couchAngles = [0 0]; % [?]
pln.propStf.numOfBeams = numel(pln.propStf.gantryAngles);
pln.propStf.isoCenter = ones(pln.propStf.numOfBeams,1) * matRad_getIsoCenter(cst,ct,0);
% dose calculation settings
pln.propDoseCalc.doseGrid.resolution.x = 5; % [mm]
pln.propDoseCalc.doseGrid.resolution.y = 5; % [mm]
pln.propDoseCalc.doseGrid.resolution.z = 5; % [mm]
% optimization settings
pln.propOpt.optimizer = 'IPOPT';
pln.propOpt.bioOptimization = 'LEMIV_effect';
%% generate steering file
stf = matRad_generateStf(ct,cst,pln);
%% dose calculation
dij = matRad_calcParticleDose(ct,stf,pln,cst);
%% inverse planning for imrt
resultGUI = matRad_fluenceOptimization(dij,cst,pln);
%% start gui for visualization of result
matRadGUI
For educational purposes, matRad is also available as a standalone application.
matRad developments were/are, in part, supported by the following research grants:
- Grant No. BA 2279/3-1 (Project No. 265744405) from the German Research Foundation (DFG)
- Grant No. WA 4707/1-1 (Project No. 443188743) from the German Research Foundation (DFG)
- Grant No. 70113094 from the German Cancer Aid